Introduction: Increased volume of epicardial adipose tissue (EAT) is now well established as an independent risk factor for all forms of atrial fibrillation (AF). EAT acts as a local compartment of immune cells and mediators that can infiltrate the myocardium given the lack of fascial boundaries between them. However, a comprehensive immune characterization of EAT is currently lacking with human tissue studies sparse and patients typically poorly matched for baseline clinical characteristics. This study sought to systematically define the immunological signature of EAT in a propensity-matched cohort of cardiac surgical patients with a prior history of AF, those in sinus rhythm and patients who developed de novo post-operative AF.
Methods: Adult patients with a history of AF and those with no prior history of AF undergoing cardiac surgery were recruited to undergo EAT, blood and subcutaneous adipose tissue (systemic and adipose tissue controls respectively) sampling. Patients were propensity-matched to ensure baseline clinical variables were similar across the groups. Immune cell isolation, flow cytometry and T lymphocyte cell stimulation assays were performed on fresh samples. Bulk RNA and spatial transcriptomic analysis were performed on a cohort of patients to determine both whole tissue and regional differences in RNA expression changes across the tissue. In 2 patients, paired EAT and right atrial appendage (RAA) tissue samples underwent combined single-cell RNA sequencing and T cell receptor sequencing.
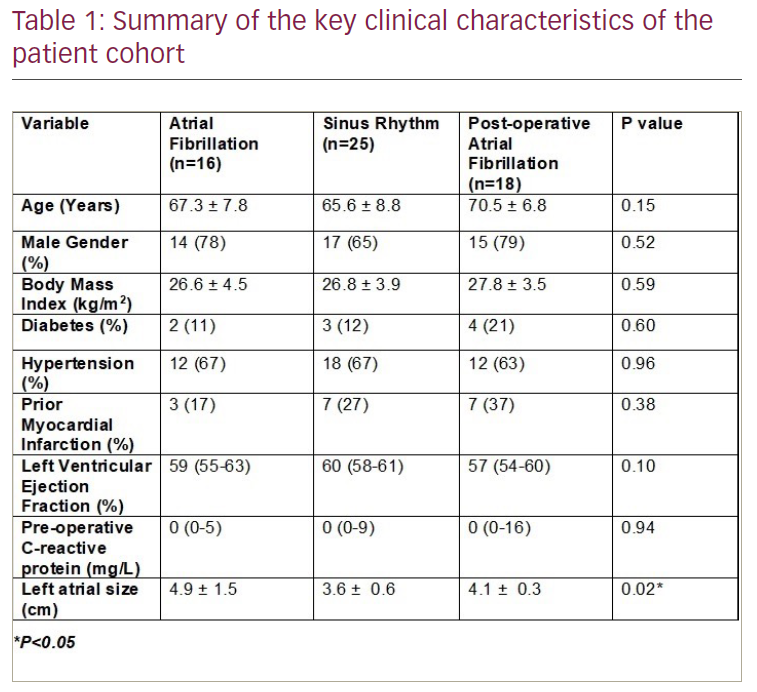
Results: A cohort of 59 propensity-matched patients was identified (Table 1). T cells were the predominant immune cell type and T-cell subset analysis in a sub-cohort of 18 patients revealed a highly significant increase in both EAT-resident CD4+ (p<0.05) and CD8+ (p<0.001) memory T-cell populations in AF patients. T-cell stimulation assays demonstrated a highly significant correlation with the proportion of tissue-resident memory (TRM) CD4+ T cells in EAT and the cytokines interferon-γ (p=0.0072) and interleukin-17 (p=0.0042). Spatial transcriptomic analysis revealed regional differences in gene expression and higher fibrosis samples exhibiting TRM-specific markers (p<0.05). In contrast, bulk EAT RNA sequencing analysis demonstrated broadly similar immune mediator expression levels between the groups. On a single-cell level, EAT and RAA exhibited comparable immune cell populations with a shared core TRM signature and TRM subsets. One of the CD8+ TRM subsets was noted to be specifically elevated in the AF cohort. Finally, T-cell receptor sequencing confirmed the same T-cell clones to be present in the RAA as the EAT.
Conclusions and implications: AF exhibits a unique EAT-resident T-cell signature that correlates with the production of the pro-inflammatory cytokines interferon-γ and interleukin-17. Single-cell RNA and T-cell receptor sequencing analysis confirms EAT to be the immune reservoir of the heart with the same T-cell clones noted to be in both EAT and RAA. EAT sampling can thus provide an accurate readout of the immune landscape of the underlying cardiac tissue.
Targeting this local resident T-cell population and the mediators it produces (e.g. using a monoclonal antibody approach) unlocks a novel paradigm in the management of the inflammatory and fibrotic components of AF genesis. ❑