Introduction: OpenEP (https://openep.io) is widely used to streamline electrophysiology research, removing the need for researchers to write code to import and analyse data. OpenEP provides a common interface for the storage, representation and analysis of electroanatomic and electrophysiological data from the major system manufacturers. Significant updates to OpenEP have been added since initial release in late 2020. This abstract describes the features and tools added recently.
Method: OpenEP is a collaborative, open-source project (https://github.com/openep) with permissive licensing. Software development has been performed in Matlab (R2020a/R2021b), Visual Studio and Python3 with a Py-QT front-end. OpenEP consists of three components: a data-parsing system; a graphical interface; and a data-analysis system. Software and system development has been performed in each of these areas.
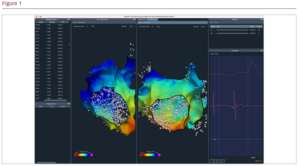
Results: Data parsing. At initial release, OpenEP supported reading in data from the Carto3 (Biosense Webster), Velocity (St Jude Medical) and Precision (Abbott) electroanatomic mapping systems. We continue to update and support the data parsers for Carto3, Velocity and Precision. In addition, we have added data parsers for the Kodex (EPD/Phillips) and EnSiteX (Abbott) mapping systems. It is also now possible to import simulation data from the openCARP simulation platform. Electroanatomic mapping data sets can also be export in the openCARP format for simulation experiments. Data analysis. Interpolation is used in electroanatomic mapping systems to create colour maps of electrical data. An extensive re-writing of code used for interpolation in OpenEP has been performed. The original generateInterpData.m function has been replaced with a new extensible architecture for data interpolation based around the openEPDataInterpolator.m class. This new architecture has uncovered previously unrecognised variation in the interpolation schemes employed in clinical mapping systems and will permit the optimisation of interpolation methods against ‘gold standard’ simulation or histological data. A similar architecture for conduction speed and velocity measurement has also been developed. This architecture exposes established techniques including the gradient, triangulation, radial basis and cosine-fit methods for calculation of conduction velocity. Conduction velocity is a key parameter defining arrhythmia mechanisms, and this tool provides a simple method for applying previously published techniques for conduction velocity calculations to clinical data. Finally, we have refined the ablation lesion quantification tools permitting time-domain analysis of radiofrequency lesion formation. Graphical interface. We are developing a graphical interface, providing the ability to visualize, manipulate and analyse electrophysiological data (see figure). To facilitate this development, we are developing a Python3-based implementation of OpenEP. A beta-testing program for the graphical interface will shortly be launched. The graphical interface provides, for the first time, a standalone system for review of electroanatomic mapping data, which can be installed on desktop/laptop computers.
Conclusion: OpenEP is an open-source platform for analysis of electro-anatomic mapping data under active development. Recent work improves data parsing and analysis functionality and facilitates usability through the development of a graphical interface.